The long-term goal of the Ramachandran Lab is to understand how distinctive chromatin landscapes that reflect cellular identity are established and maintained.
Regulation of genome access underlies growth, development, differentiation, and disease states including tumor initiation. Genome access is regulated by dynamic chromatin landscapes. We want to uncover fundamental mechanisms that shape chromatin landscapes by mapping chromatin structure at high temporal and spatial resolution using new experimental and/or computational methods. This research program in chromatin biology is at the intersection of genomics, biochemistry, and structural biology.
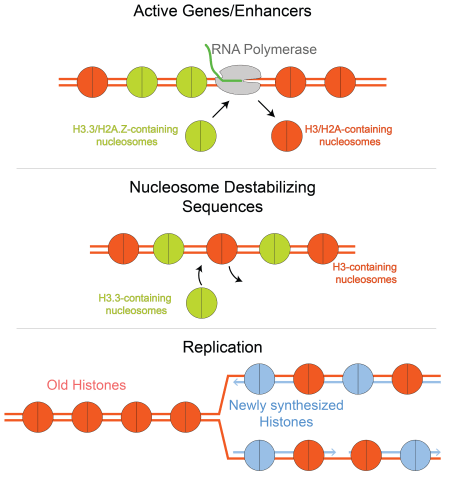
Chromatin landscape, the nature of proteins bound at different locations in the chromosome, emerges from competing activities of nucleosome (re)assembly, transcription factor binding, and chromatin remodeler activity overlaid on metabolic processes that occur on the chromosome like transcription, replication, and repair. To understand how chromatin landscapes are established and maintained, we need to uncover the basis for biochemical activity on a chromatinized genome. We are uncovering this by answering two fundamental questions:
- What determines the subset of sites on the genome bound by sequence-specific DNA-binding proteins (also called transcription factors)?
- How are repressed chromatin states maintained through successive cell divisions?
In parallel, we are developing new genomic methods to map structural properties of chromatin in cells.
Some of our earlier work: